Thank you for your interest in Montana Molecular’s assays for SARS-CoV-2 research and drug discovery. In a rapidly evolving field, as new insights into the mechanisms of viral entry come to light, we sincerely appreciate your patience and feedback as we continue to optimize these assays.
We are continuing development of BacMam-based tools, including pseudoviruses bearing the SARS-CoV-2 Spike protein, and fluorescent reporters to enable host cell entry. BacMam is based on a BSL-1 baculovirus that is non pathogenic in humans, and does not replicate in mammalian cells, or integrate into the genome.
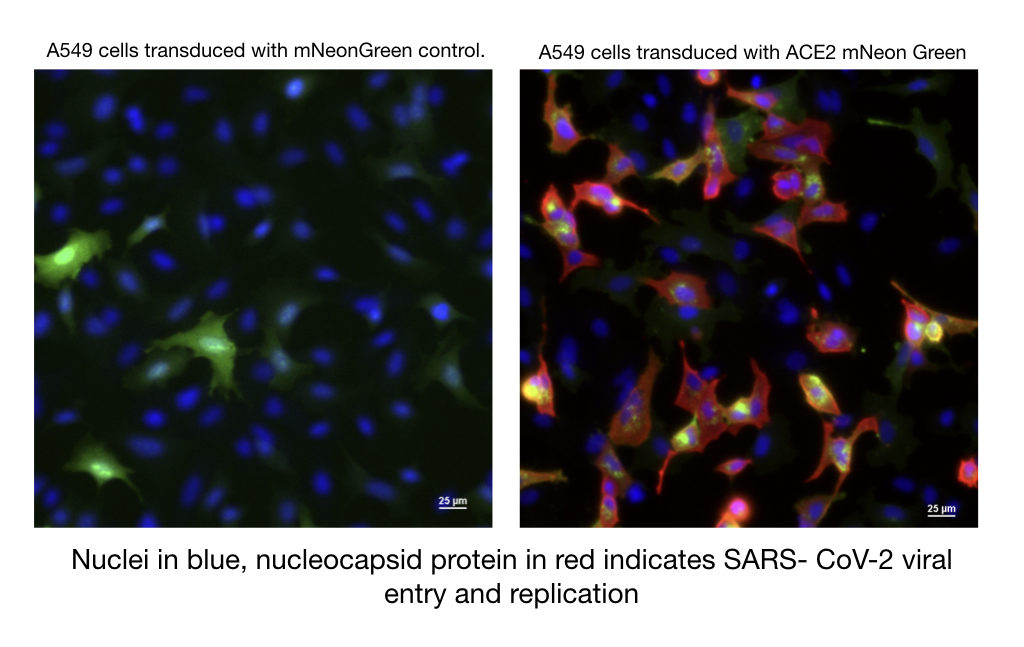
BacMam expression kits carrying the host receptor ACE-2 are available with either a red or green fluorescent protein attached. These produce bright fluorescence that can be read on a plate reader or imaging system. Following reports that ACE2 and TMPRSS2 co-expression is sufficient for viral entry (Hoffmann et al. 2020), we packaged TMPRSS2, in BacMam. This host protease is included in our ACE-2 kits. An early adopter working in a BSL-3 facility showed that A549 cells expressing our green ACE-2 protein become permissive to infection with the real SARS-CoV-2 virus.
To avoid the need to screen under BSL-3 conditions, we are also developing a series of pseudoviruses, based on BacMam. Initially, we replaced the BacMam VSVG pseudotype with full length Spike protein. This pseudo-SARS-CoV-2 expresses either green or red fluorescent proteins that are targeted to the nucleus of the host cell, so that viral entry is easily detectable. Last weekend, we tested a “Bald” BacMam virus without VSVG or Spike. This control revealed that baseline transduction is driven purely by the GP64 envelope glycoprotein inherent in BacMam. By matching the number of viral genes used in the Spiked pseudovirus and the “Bald” control virus, Spike-driven viral entry can be determined by the relative amount of fluorescence above the baseline established by the control. Early adopters who have already received the pseudo-SARS-CoV-2 can request a free sample of the bald BacMam control.
These experiments with pseudovirus in HEK293 and A549 cells indicate that additional host components may also be necessary, at least in these cell types. There is growing evidence that trypsin treatment increases Spike driven transduction (Menachery et al. 2020; Zheng et al. 2018), as well as Cathepsin L (Huang et al. 2006; Ou et al. 2020). We are currently testing each of these treatments to determine their effect on the pseudovirus signal in HEK293 and A549 cells. We’ll have an update by early next week and appreciate your patience as the information about SARS-CoV-2 entry is in flux. Please see the list of references for more details.
In addition, a new series of pseudo-SARS-CoV-2 designs are in production to improve Spike presentation. In one case, the C-terminus of the Spike protein has been replaced with GP64, a strategy known to improve the amount of pseudotype protein on the BacMam surface. We’ll be harvesting BacMam and testing this next generation of pseudo-SARS-CoV-2 over the next week and will have another update June 25. Early adopters who have already received the first generation pseudovirus kit will receive a free upgrade to the new pseudovirus.
A pseudovirus carrying the D614G Spike which originally appeared in a European isolate is also in production. This mutation appears to have functional consequences for viral entry (Korber et al. 2020; Zhang et al. 2020). QA/QC for the D614G mutant is scheduled for mid July.
Huang, I-Chueh, Berend Jan Bosch, Fang Li, Wenhui Li, Kyoung Hoa Lee, Sorina Ghiran, Natalya Vasilieva, et al. 2006. “SARS Coronavirus, but Not Human Coronavirus NL63, Utilizes Cathepsin L to Infect ACE2-Expressing Cells.” The Journal of Biological Chemistry 281 (6): 3198–3203.
Korber, B., W. M. Fischer, S. Gnanakaran, H. Yoon, J. Theiler, W. Abfalterer, B. Foley, et al. 2020. “Spike Mutation Pipeline Reveals the Emergence of a More Transmissible Form of SARS-CoV-2.” bioRxiv. https://doi.org/10.1101/2020.04.29.069054.
Menachery, Vineet D., Kenneth H. Dinnon 3rd, Boyd L. Yount Jr, Eileen T. McAnarney, Lisa E. Gralinski, Andrew Hale, Rachel L. Graham, et al. 2020. “Trypsin Treatment Unlocks Barrier for Zoonotic Bat Coronavirus Infection.” Journal of Virology 94 (5). https://doi.org/10.1128/JVI.01774-19.
Ou, Xiuyuan, Yan Liu, Xiaobo Lei, Pei Li, Dan Mi, Lili Ren, Li Guo, et al. 2020. “Characterization of Spike Glycoprotein of SARS-CoV-2 on Virus Entry and Its Immune Cross-Reactivity with SARS-CoV.” Nature Communications 11 (1): 1620.
Zhang, Lizhou, Cody B. Jackson, Huihui Mou, Amrita Ojha, Erumbi S. Rangarajan, Tina Izard, Michael Farzan, and Hyeryun Choe. 2020. “The D614G Mutation in the SARS-CoV-2 Spike Protein Reduces S1 Shedding and Increases Infectivity.” bioRxiv. https://doi.org/10.1101/2020.06.12.148726.
Zheng, Yuan, Jian Shang, Yang Yang, Chang Liu, Yushun Wan, Qibin Geng, Michelle Wang, Ralph Baric, and Fang Li. 2018. “Lysosomal Proteases Are a Determinant of Coronavirus Tropism.” Journal of Virology 92 (24). https://doi.org/10.1128/JVI.01504-18.
