Quick-View Protocol
About These Assays #D0201G Green Cilia-targeted and #D0211G Ratiometric Cilia-targeted cADDis cAMP
Cyclic AMP is an essential second messenger that may regulate pathways inside primary cilia. To measure cAMP within the ciliary space, we fused cADDis, a genetically encoded sensor for cAMP, to the 5HT6 receptor. The cilia-targeted cADDis sensor decreases in fluorescence intensity when localized cAMP is increasing in the cilia. The following protocol assumes cilia formation in your cells (Delling 2013).
Overview
The green cADDis sensor (Tewson, 2015) is a fluorescent protein-based sensor for live cell measurement of cAMP in the primary cilium of mammalian cells. The vector carrying this sensor is BacMam, a modified baculovirus, which can be used for delivery to a wide variety of mammalian cell types, including many primary cultures.
In mammalian cells, BacMam expresses only the fluorescent sensor and is a BSL-1 reagent. We recommend that you take the time to do a dilution series of BacMam in your favorite cells, to optimize efficiency in your particular cells and experiments.
Materials in the Kit
Green Cilia-Targeted cADDis cAMP sensor BacMam in TNM-FH Insect Culture Medium (Allele Biotech product #ABP-MED-10001).
-Green fluorescent sensor that decreases in fluorescence intensity in response to increases in cAMP.
-#D0201G is the green sensor only, #D0211G is labeled with a red fluorescent tag for ratiometric measurement.
Sodium Butyrate (Sigma Aldrich product # B5887) 500 mM in H2O.
-Sodium Butyrate is added to the culture to maintain BacMam expression. Other HDAC inhibitors may work as well.
Additional Materials Not Supplied
- Greiner CellCoat (#655946) is our preferred 96-well plate available from VWR.
- Dulbecco’s Phosphate Buffered Saline (DPBS) available from VWR [Dulbecco, R. and Vogt, M.1957].
- Complete culture media specific to your cells. (Please see Assay Performance Section)
- 2-methyl-5hydroxytryptamine (2-M-5HT) a 5HT3 receptor agonist
- L-858051 a forskolin derivative
Storage/Biosafety/Warranty
BacMam stocks should be stored at 4°C and protected from light. Avoid repeated freeze/thaw cycles.
BioSafety Considerations
BacMam does not replicate in mammalian cells and expresses only the fluorescent sensor. While it should be handled carefully, in a sterile environment, it is classified as a Biosafety Level 1 (BSL-1) reagent.
This product is for research use only and is not recommended for use or sale in human or animal diagnostic or therapeutic products.
Warranty
Materials are provided without warranty, express or implied. End user is responsible for making sure product use complies with applicable regulations. No right to resell products or any components of these products is conveyed.
DAY 1: Transduce and Plate Cells
This protocol is optimized for rapidly dividing, immortalized cell lines. However, the protocol can be adjusted for transducing non-dividing adherent cells such as neurons, islets, cardiomyocytes, and iPSC-derived lines. We recommend that you take the time to optimize the assay for your particular cell type. See our Suggestions for Adherent Cells following this protocol.
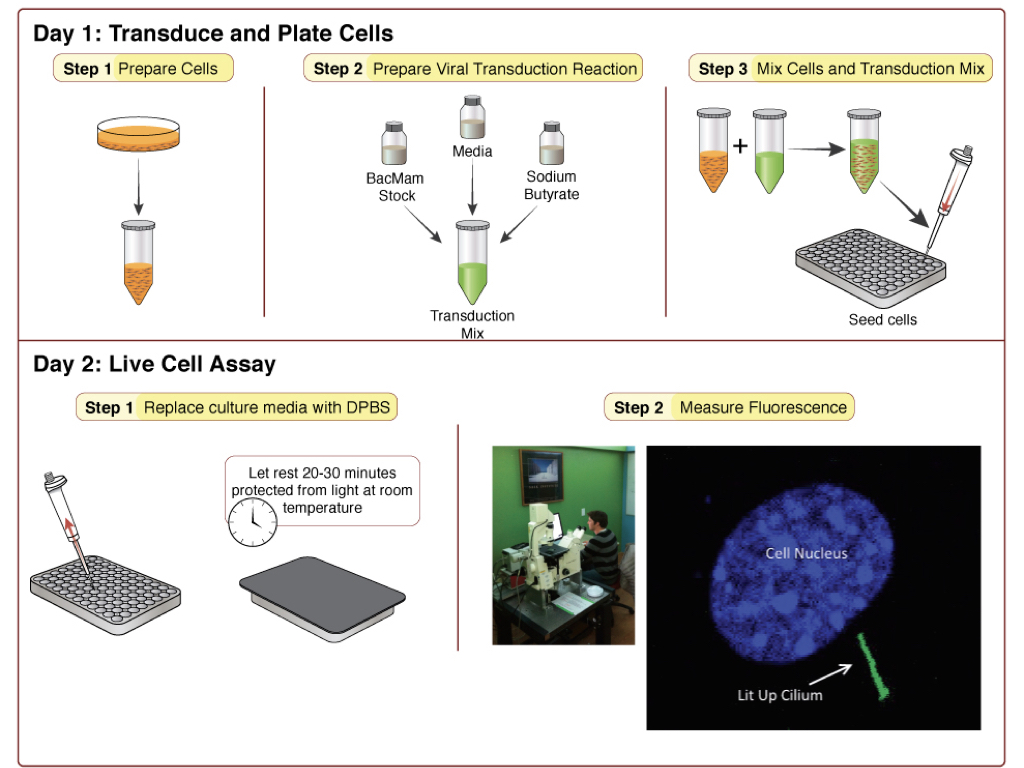
Step 1: Prepare cells (Tube A)
- Detach cells from flask using normal trypsinization protocol. Resuspend cells in complete culture media and determine cell count.
- Prepare a dilution of cells at your desired concentration*. 100 μl of this cell resuspension will be required for a single well in a 96-well plate, so prepare enough of the dilution to seed the desired number of wells in the plate. Let cells sit in hood and move on to preparation of the viral transduction reaction.
*450,000 cells/mL works well for HEK293 cells.
Step 2: Prepare Viral Transduction Reaction (Tube B)
- Prepare a 500 mM stock solution of sodium butyrate in sterile water (in your kit).
- For each transduction reaction (i.e. one well in a 96-well plate), prepare the transduction solution by mixing 25 μl of the BacMam sensor stock with 0.6 μl of the 500 mM stock solution of sodium butyrate** and 24.4 μl of the complete culture media for your cells, for a total volume of 50 μl. Mix gently.
** Concentration of sodium butyrate should be 6mM in this step. Following Step 3, final concentration of sodium butyrate will be 2mM.
Step 3: Mix Cells and Transduction Mix from above.
- Mix Tube A and Tube B (100 μl tube A + 50 μl tube B). Mix gently and then seed 150 μl of mix per well on the 96-well plate.
- Cover plate with aluminum foil to protect from light and incubate at room temperature for 30 minutes.
- Incubate overnight under normal growth conditions, protected from light.
DAY 2: Measuring Fluorescence
Cells are now ready for assay. Prior to imaging, replace culture media with DPBS. Wash gently so as not to dislodge cells. Cover the cells and allow them to rest at room temperature in DPBS for 20-30 minutes before measuring fluorescence. Experiments are performed at 25°C using standard GFP excitation and emission wavelengths. Figure 1 shows an example of the sensor response in gray in IMCD3 cells, imaged on the left (Moore 2016).
Fluorescence Properties
cADDis is constructed with the very bright, mNeon green fluorescent protein. We recommend Chroma’s Catalog set #49003 for optimal results. Excitation max: 506nm Emission max: 517nm
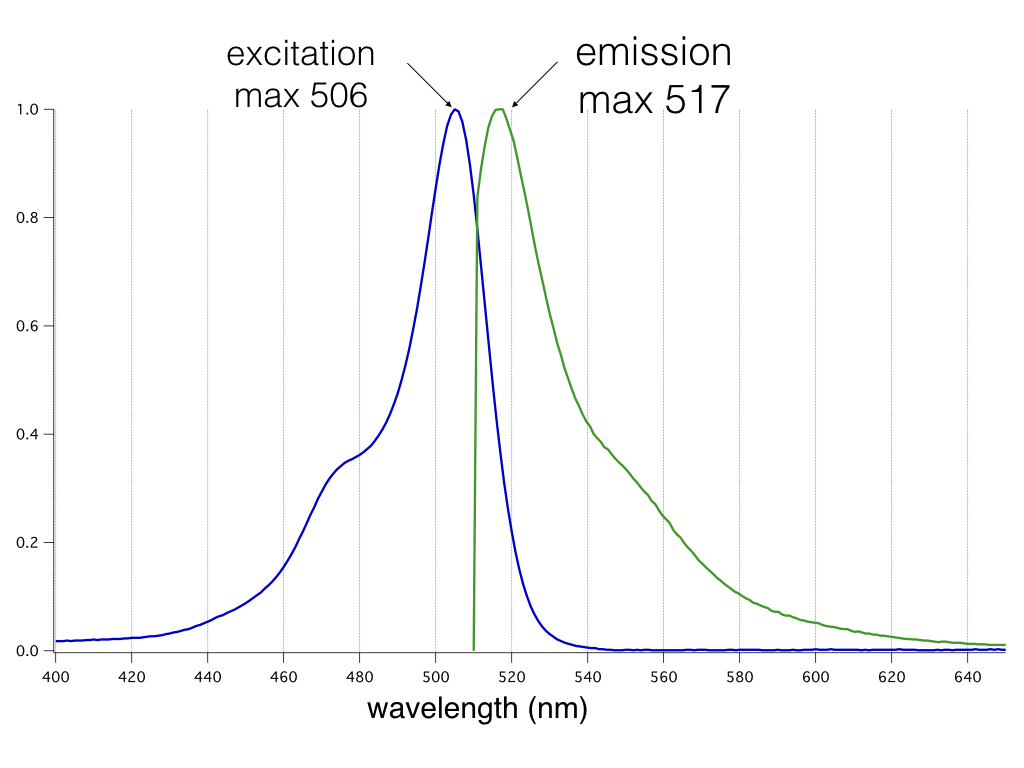
Absorption and emission properties of the mNeon green fluorescent protein plotted as a function of wavelength.
Assay Performance Considerations
Expression levels of the sensor.
To optimize the assay in your particular cell type, it is important to optimize the amount of virus used in the transduction. Too little virus will produce variable results particularly if the sensor expression levels are low and difficult to detect on your instrument.
Receptor expression
The magnitude of the sensor response can be affected by the level of GPCR expression in your cells. We have found that low levels of receptor expression produce the largest signals, while high levels of receptor expression often produce smaller responses. This is consistent with the observation that over expression of some GPCRs can change the levels of second messengers due to low levels of spontaneous activity.
Culture Media Considerations
This assay has been validated with EMEM, DMEM, and F-12K complete growth media. Other types of media may affect results.
Contact Us
If you have ideas about how we can improve our products or protocols, then we would like to hear from you. Your feedback is extremely valuable. Please send an email to info@montanamolecular.com. We’ll respond as quickly as we can.
References
1. Tewson PH, Martinka S, Shaner N, Hughes TE, Quinn AM: New DAG and cAMP sensors optimized for live cell assays in automated laboratories. Journal of Biomolecular Screening 2015.
2. Delling M, DeCaen PG, Doerner JF, Febvay S., Clapham DE, Primary cilia are specialized calcium signalling organelles. Nature. 504, 311-314 (2013).
3. Borner S, Schwede F, Schlipp A, Berisha F, Calebiro D, et al. (2011) FRET measurements of intracellular cAMP concentrations and cAMP analog permeability in intact cells. Nat Protoc 6(4): 427-438. 10.1038/nprot.2010.198 [doi].
4. Moore, B.S. et.al. Cilia Have High cAMP Levels That Are Inhibited by Sonic Hedgehog Regulated Calcium Dynamics. PNAS 2016, vol. 113 no. 46. doi: 10.1073/pnas.1602393113
5. Shaner, N.C., Lambert, G.G., Chammas, A., Ni, Y., Cranfill, P.J., Baird, M.A., Sell, B.R., Allen, J.R., Day, R.N., Israelsson, M., Davidson, M.W., & Wang, J. (2013) “A bright monomeric green fluorescent protein derived from Branchiostoma lanceolatum.” Nature Methods, May;10(5):407-9. doi: 10.1038/nmeth.2413.



