SARS CoV-2 Pseudoviruses
Tools for Measuring SARS-CoV-2
SARS-CoV-2 Pseudovirus
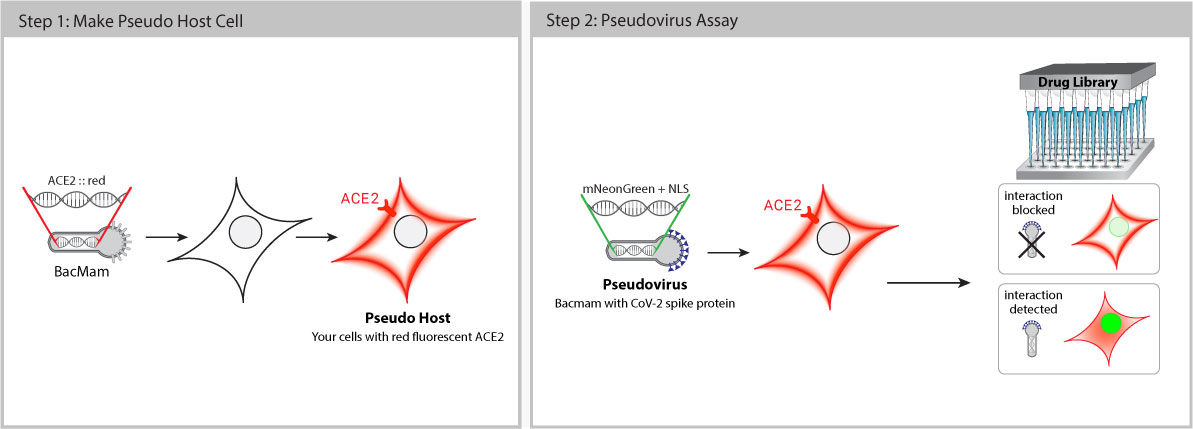
Pseudo SARS-CoV-2 is safe to use with no risk of infection. These pseudoviruses are BSL-1 baculoviruses pseudotyped with Spike (S) proteins. This means that biologists and drug discovery teams can work on COVID-19 projects outside of BSL-3 facilities with standard safety procedures. Baculovirus production in Sf9 cells is cost effective with low batch-to-batch variability.
Once inside the cell, the pseudovirus does not replicate, but delivers a genetically-encoded, fluorescent reporter that expresses bright green fluorescence in the host cell nucleus. When Spike-ACE2 interaction is successfully blocked, the host cell fluorescence decreases.
Three versions of Pseudo SARS-CoV-2 are available. One (#C1110G) carries the Spike protein originally sequenced from isolates obtained in Wuhan, and another (#C1120G) carries D614G Spike (Korber et.al. 2020). #C1123G carries the spike protein found in the Delta Variant, or B.1.617.2. These constructs have been sequence verified and tested in A549 cells. Pseudovirus affinity for the host receptor ACE2 was confirmed using absorption experiments on A549 cells expressing different amounts of ACE2. Further validation in other cell types is ongoing.
As SARS-CoV-2 evolves, researchers are working to understand how Spike mutations affect transmission and their potential effect on the efficacy of blocking antibodies. New pseudoviruses reflecting emerging mutations and variants are continually in development, we would love to hear what versions would be useful to you! Let us know at info@montanamolecular.com.
Order a SARS CoV-2 Pseudovirus Kit
Viral Entry Assay in Primary Human Endothelial Cells
Assay measures SARS-CoV-2 spike protein / ACE2 receptor dependent entry into endothelial cells. HUVECs were transduced with ACE2 receptor, then infected with baculovirus pseudotyped with SARS-CoV-2 spike protein. Pretreatment with the known entry inhibitor, lactoferrin, (Mirabelli et al., 2020) showed dose-dependent inhibition.
Pseudovirus Absorption Assay
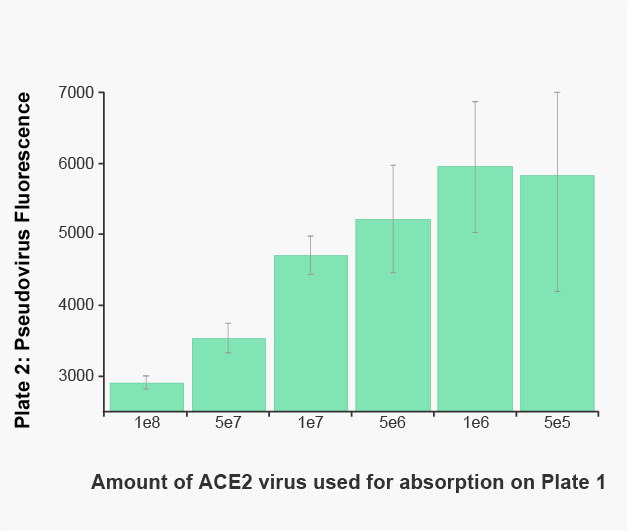
A549 cells in a 96-well plate received increasing amounts of red fluorescent ACE2, or a red fluorescent protein control. The following day, Pseudo SARS-CoV-2 pseudovirus was added to the plate and incubated for 2 hours. The media from the plate was transferred to a second plate of fresh A549 cells expressing a constant amount of ACE2. Twenty-four hours later, the cells were washed with PBS and the green fluorescence produced by the pseudovirus from the media was measured on a BioTek Synergy™ plate reader. The two hour incubation on ACE2 expressing cells clearly reduces the pseudovirus concentration in an ACE2 dependent manner.
Blocking Pseudovirus
Pseudo SARS-CoV-2 is an enveloped virus that presents the full length spike protein on the viral surface membrane with the same topology as SARS-CoV-2. The spike protein is oriented perpendicular to the membrane such that multiple antibody binding sites are available and presented in a structurally relevant way. Two versions are available, one with the sequence identified in the original Wuhan reference specimen and the newer D614G mutant that first appeared in Europe and is rapidly replacing the Wuhan strain.
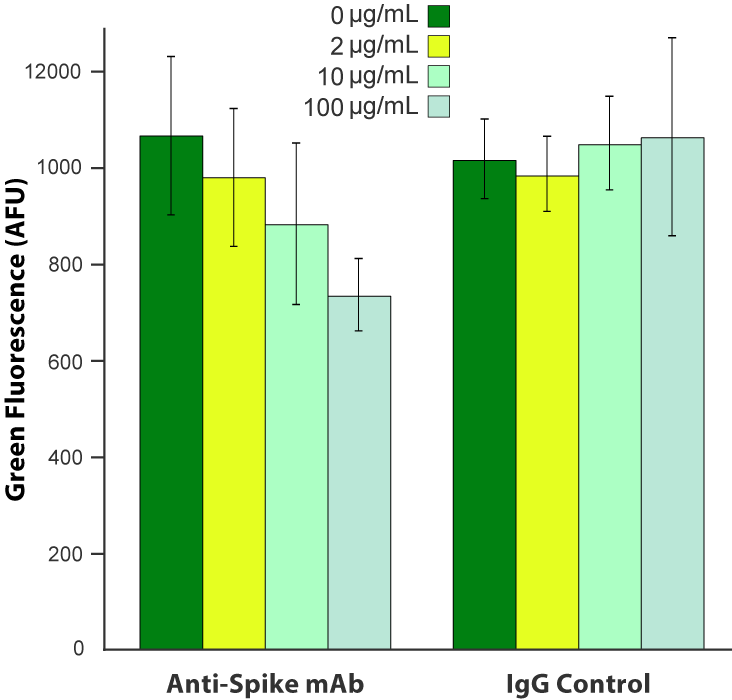
SARS-CoV-2 pseudovirus is blocked by anti-Spike antibody. A549 cells were transduced with 6.6 x 10 8 VG/mL Fluorescent ACE2 (#C1100R). The following day, cells were treated with a monoclonal anti-Spike antibody (Acro Biosystems, catalog # SAD-S35) or an equivalent amount of IgG isotype control, then challenged with 3.3 x 10 8 VG/mL SARS-CoV-2 pseudovirus (#C1110G). Green fluorescence was quantified ~20 hours later.
Korber, Bette, Will M. Fischer, Sandrasegaram Gnanakaran, Hyejin Yoon, James Theiler, Werner Abfalterer, Nick Hengartner, et al. 2020. “Tracking Changes in SARS-CoV-2 Spike: Evidence That D614G Increases Infectivity of the COVID-19 Virus.” Cell, July. https://doi.org/10.1016/j.cell.2020.06.043.
Contact Us
If you have any questions about using these tools in your cells or would like to request a quote, send us an email at info@montanamolecular.com.
For updates on COVID-19 tools for research and drug discovery, visit our NEWS page.